
Research Techniques Made Simple: Molecular Docking in Dermatology - A Foray into In Silico Drug Discovery - ScienceDirect
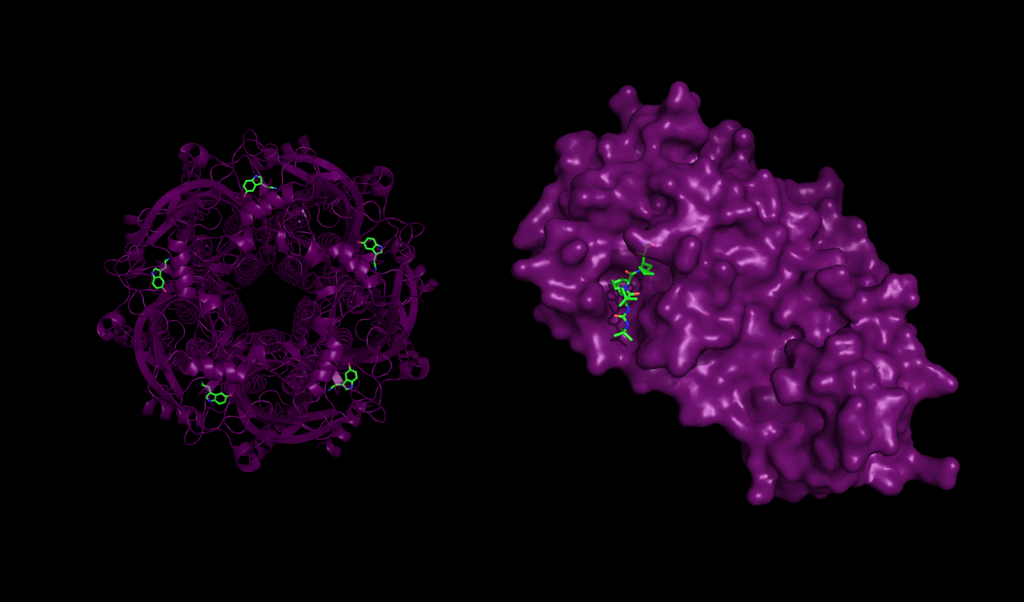
Docking Proteins to Deny Disease: Computational Considerations for Simulating Protein-Ligand Interaction | Exxact Blog

Molecular Dynamics in Mixed Solvents Reveals Protein–Ligand Interactions, Improves Docking, and Allows Accurate Binding Free Energy Predictions | Journal of Chemical Information and Modeling
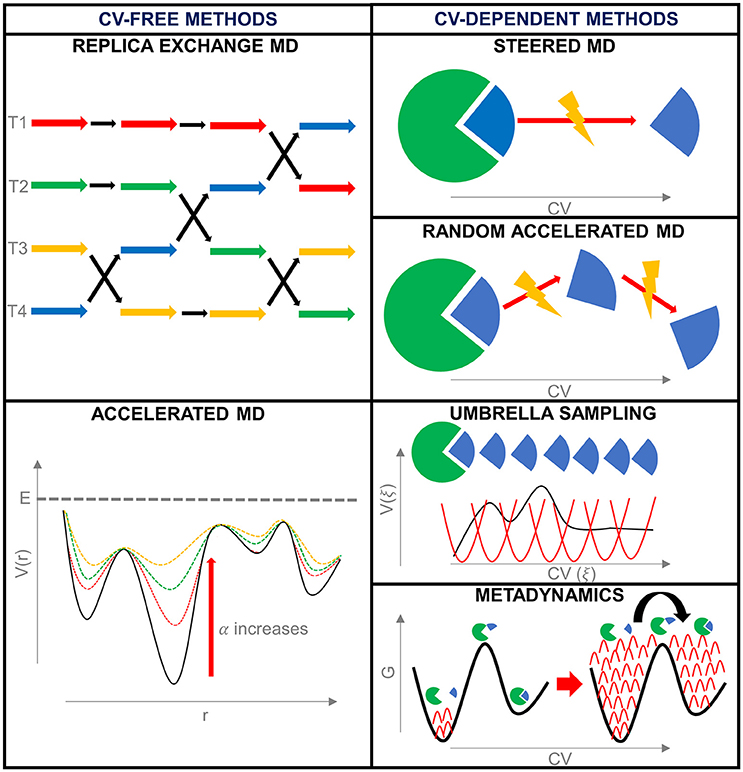
Frontiers | Bridging Molecular Docking to Molecular Dynamics in Exploring Ligand-Protein Recognition Process: An Overview

Research Techniques Made Simple: Molecular Docking in Dermatology - A Foray into In Silico Drug Discovery - ScienceDirect

The In Silico Fischer Lock-and-Key Model: The Combined Use of Molecular Descriptors and Docking Poses for the Repurposing of Old Drugs | SpringerLink

Schematic illustrations of the three protein-ligand binding models: (a)... | Download Scientific Diagram
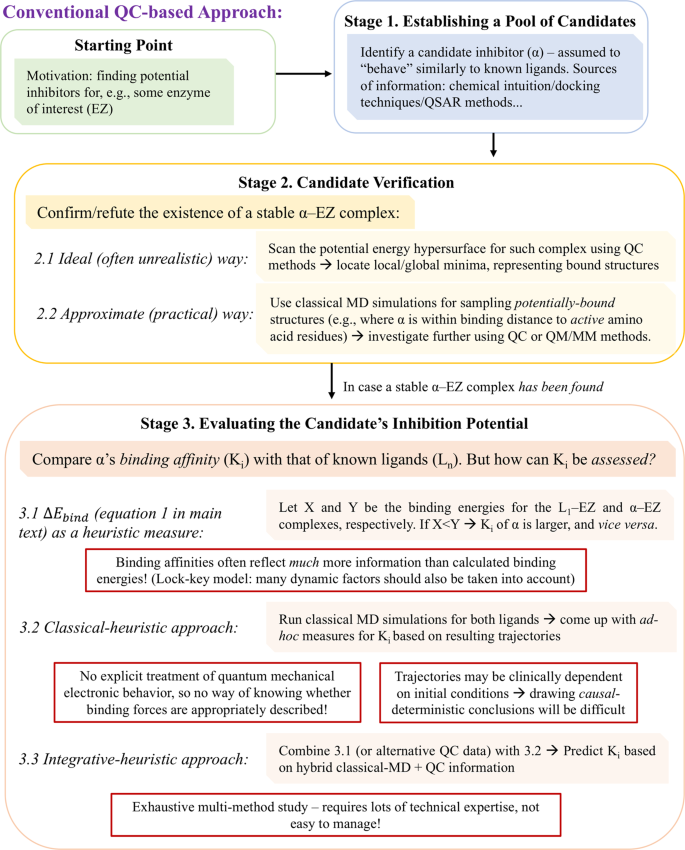

Toward Simple, Predictive Understanding of Protein-Ligand Interactions: Electronic Structure Calculations on Torpedo Californica Acetylcholinesterase Join Forces with the Chemist's Intuition | Scientific Reports
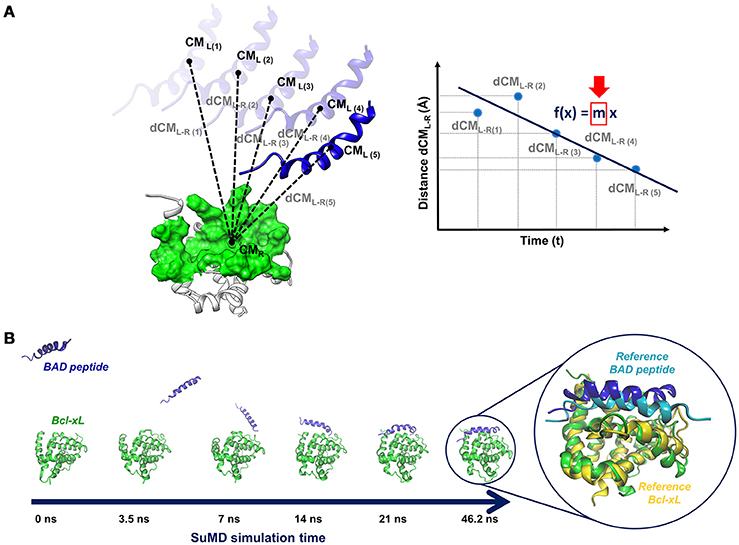
Frontiers | Bridging Molecular Docking to Molecular Dynamics in Exploring Ligand-Protein Recognition Process: An Overview

Different technique for Molecular docking: a) Induced work docking b)... | Download Scientific Diagram
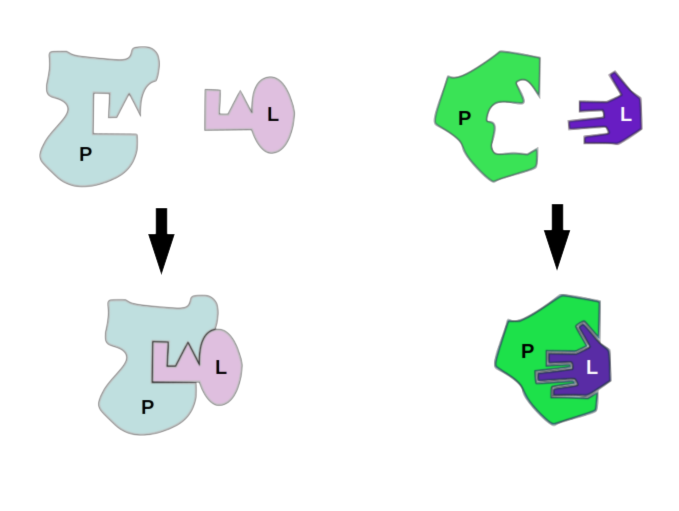
Docking Proteins to Deny Disease: Computational Considerations for Simulating Protein-Ligand Interaction | Exxact Blog






![PDF] Molecular Docking: From Lock and Key to Combination Lock. | Semantic Scholar PDF] Molecular Docking: From Lock and Key to Combination Lock. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/440d47cf86b98dc01689c8fb3b49815334782c3e/2-Figure1-1.png)


![PDF] Molecular Docking: From Lock and Key to Combination Lock. | Semantic Scholar PDF] Molecular Docking: From Lock and Key to Combination Lock. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/440d47cf86b98dc01689c8fb3b49815334782c3e/2-Figure2-1.png)
![PDF] Molecular Docking: From Lock and Key to Combination Lock. | Semantic Scholar PDF] Molecular Docking: From Lock and Key to Combination Lock. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/440d47cf86b98dc01689c8fb3b49815334782c3e/6-Figure3-1.png)
